02/25/2008
|
Glimpses into the mechanisms of RNA catalysis revealed by molecular simulations
Recent research from the group of Professor Darrin York.
Cover Story in Chemistry & Biology, from the group of Prof. William
Scott (UC Santa Cruz) in collaboration with Prof. Darrin York (UMN) "Solvent
Structure and Ribozyme Catalysis"; Vol 15, 332-342, 21
April 2008 (Link) |
 |
An area of intense
experimental and theoretical research effort has been concentrated on
elucidating how RNA molecules are able to catalyze complex biochemical
reactions. A detailed understanding of the underlying mechanisms of these RNA
enzymes provides insight into the inner workings of more complex cellular
catalytic machinery such as the ribosome. Ultimately, these insights may aid
the rational design of new medical therapies that target viral, neurological
and genetic disease, as well as the development of new bio/nanotechnology.
Recently, the research group of Prof. Darrin
York, including research associate Dr. Tai-Sung Lee, MSI research scholar Dr. Carlos
Silva-Lopez and graduate student George Giambasu of the Department of
Chemistry, in collaboration with Prof. Bill Scott
of the Department of Chemistry, UCSC, have mapped out a plausible mechanism
for the role of divalent metal ions in hammerhead ribozyme catalysis from
molecular dynamics simulations (Fig. 1). Complementary work, done in
collaboration with Prof. Jiali
Gao of the Department of Chemistry and Dr. Kwangho Nam of the Department
of Chemistry and Chemical Biology at Harvard University, on the hairpin
ribozyme, suggest
the electrostatic environment of the hairpin ribozyme accounts for the vast
majority of its catalytic proficiency without recruitment of divalent metal
ions involved in the chemical steps of
the reaction (Fig. 2). Together, these studies provide two-fold insight into
how molecules of RNA are able to catalyze biochemical reactions with and
without chemical participation by divalent metal ions in the active site. The
results of these two studies are currently in press in the Journal of the American Chemical Society.
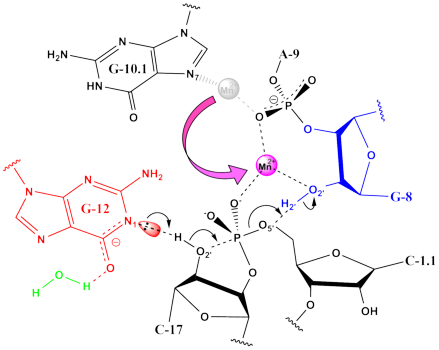
One metal
mechanism in hammerhead ribozyme derived from simulations of the reactant,
early and late transition states.
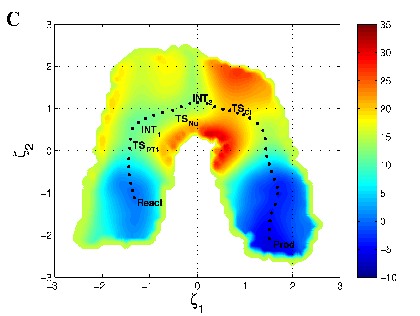
Two-dimensional
free energy profile for hairpin ribozyme calculated with a new AM1/d-PhoT model
for phosphoryl transfer.
|