|
01/26/2010
Simulations of an RNA Enzyme Molecular Switch
Recent research from the research group of Professor Darrin York.
Recently, graduate student George Giambasu, research associate Dr.
Tai-Sung Lee and Prof. Darrin York of the Department of Chemistry, along
with collaborators Carlos P. Sosa from IBM and William Scott and Michael
Robertson of the Center for Molecular Biology of RNA at UCSC have used
large scale molecular dynamics simulations to reveal the dynamical hinge
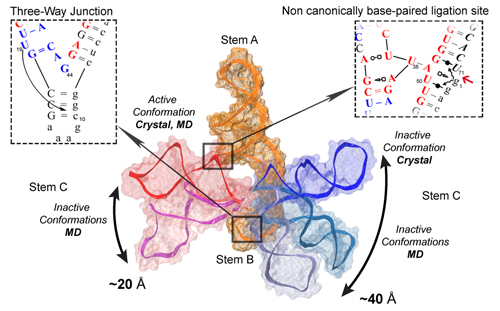
points of the L1 ligase all-RNA synthetic molecular switch. The L1
ligase ribozyme is an in vitro selected ribozyme that uses a
non-canonically base-paired ligation site to catalyze regioselectively
and regiospecifically the 5' to 3' phosphodiester bond ligation, a
reaction relevant to origin of life hypotheses that invoke an RNA World
scenario. From these simulations, along with crystallographic data, a
set of 4 virtual torsion angles that span two evolutionarily conserved
and restricted regions were identified as dynamical hinge points in the
conformational switch transition. The ligation site visits three
distinct states characterized by hydrogen bond patterns that are
correlated with the formation of specific contacts that may promote
catalysis. The insights gained from these simulations contribute to a
more detailed understanding of the coupled catalytic/conformational
switch mechanism of L1 ligase that may facilitate the design and
engineering of new catalytic riboswitches.
This work is in press in the RNA Journal.
|