Main navigation | Main content
10/06/2010
Recent research from the research group of Professor Jiali Gao
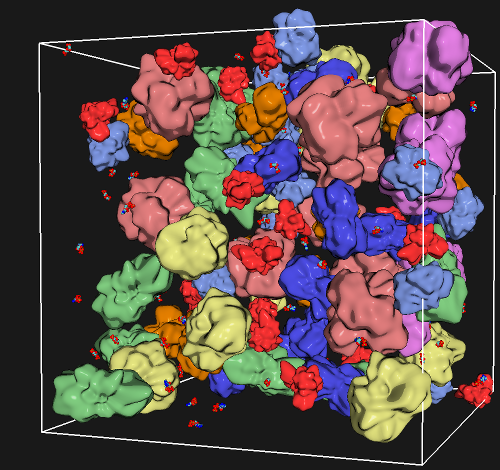
The next challenge in computational biology is to model the dynamics and signaling processes taking place in a biological cell. To achieve this goal, macromolecular particles in the cell such as proteins, nucleic acids and the ribosome must be represented by coarse-grained approaches. Graduate student Michael Mazack of the Scientific Computation Graduate Program and Professor Jiali Gao recently developed an analytically coarse-grained method (ACG), in which a macromolecular structure is treated as an entity of uniform mass density without the explicit atomic details; yet, the individual macromolecular species still retain the information of, and are constructed based on the detailed atomistic coordinates determined experimentally. The excluded volume and surface of the ACG macromolecule are explicitly represented by a spherical harmonic basis, which can provide any desired accuracy and detail depending on the problem of interest. Importantly, physical and biological properties can be described by exactly the same mathematical procedure as the representation of the macromolecule. The treatment of the internal dynamics of an ACG protein was recently featured in the Journal of Chemical Theory and Computation (Link: http://pubs.acs.org/doi/abs/10.1021/ct100426m), which paves a way to use physics-based simulations to study macromolecular interactions and assembly in biological cells.