Main navigation | Main content
10/01/2012
Recent research from the research group of Professor
An article by scientists from the Department of Chemistry's Chemical Theory Center and the University of Minnesota's Supercomputing Institute is one of the most downloaded articles in the last 90 days in Chemical Physics.
The article, "A Benchmark Test Suite for Proton Transfer Energies and Its Use to Test Electronic Structure Model Chemistries," reflects the research of Santhanamoorthi Nachimuthu, Ph.D., a former postdoctoral research fellow
in Professor Donald Truhlar's group who is now at the Institute of
Atomic and Molecular Sciences in the Academia Sinica in Taiwan, Professor Jiali Gao, and Truhlar. Their article was published in May 2012, and is now the fourth most downloaded article.
Abstract
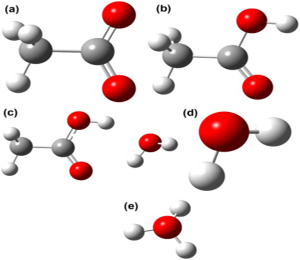
We present benchmark calculations of nine selected points on potential energy surfaces describing proton transfer processes in three model systems, View the MathML source, H5O2+, CH3OH...H+...OH2, and CH3COOH...OH2. The calculated relative energies of these geometries are compared to those calculated by various wave function and density functional methods, including the polarized molecular orbital (PMO) model recently developed in our research group and other semiempirical molecular orbital methods. We found that the SCC-DFTB and PMO methods (the latter available so far only for molecules consisting of only O and H and therefore only for the first of the three model systems) give results that are, on average, within 2 kcal/mol of the benchmark results. Other semiempirical molecular orbital methods have mean unsigned errors (MUEs) of 3–8 kcal/mol, local density functionals have MUEs in the range 0.7–3.7 kcal/mol, and hybrid density functionals have MUEs of only 0.3–1.0 kcal/mol, with the best density functional performance obtained by hybrid meta-GGAs, especially M06 and PW6B95.
To read the complete article, go to Chemical Physics.
Figure: The structures of (a) CH<ce:inf loc="post"> 3</ce:inf> COO<ce:sup loc="post"> ?</ce:sup> . (b) CH<ce:inf loc="post"> 3</ce:inf> COOH, (c) CH<ce:inf loc="post"> 3</ce:inf> COOH?H<ce:inf loc="post"> 2</ce:inf> O, (d) H<ce:inf loc="post"> 2</.